A 2025 review subject was non-electrophilic Nrf2 activators:
“NRF2 can be induced via:
- Non-specific electrophile/ROS generation,
- Disruption of the NRF2–KEAP1 protein–protein interaction,
- Autophagy-mediated KEAP1 degradation,
- Direct modulation of NRF2 protein stability, and
- Post-transcriptional/post-translational modifications.
Except for a single intervention, therapeutic hypothermia, every non-pharmacological strategy with defined mechanisms employs more than one of these routes, most frequently pairing post-translational modification with either protein-stability regulation or limited electrophile production. This combinatorial activation elevates both NRF2 abundance and transcriptional competence while minimizing the liabilities of purely electrophilic agents and circumventing the efficacy limitations.
Classical electrophilic NRF2 activators, despite potent activation potential, exhibit paradoxically reduced therapeutic efficacy relative to single antioxidants, attributable to concurrent oxidative stress generation, glutathione depletion, mitochondrial impairment, and systemic toxicity. Although emerging non-electrophilic pharmacological activators offer therapeutic potential, their utility remains limited by bioavailability and suboptimal potency.”
https://www.mdpi.com/2076-3921/14/9/1047 “Non-Electrophilic Activation of NRF2 in Neurological Disorders: Therapeutic Promise of Non-Pharmacological Strategies”
These researchers exaggerated problems of electrophilic Nrf2 activators such as “mitochondrial impairment, and systemic toxicity” so they could have something to write about. Just like every intervention, the dose determines the response. I can’t imagine not eating broccoli sprouts in favor of brain zapping with electroconvulsive therapy or transcranial magnetic stimulation just to avoid sulforaphane’s temporary mild oxidative stress that activates Nrf2 for 15-20 minutes.
But there are limitations to how an unwell person can benefit from Nrf2 activation. For example, I haven’t curated many cancer papers because healthy body functioning can’t be assumed.
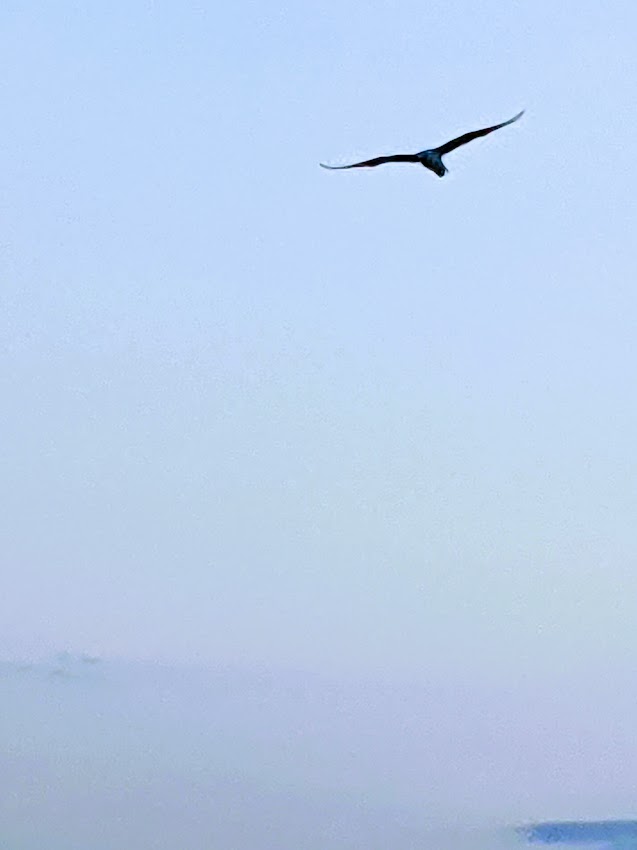
I walk the beach at sunrise, weather permitting, because it makes me feel good, and I’m always happy afterwards that I made the effort to get outside. That doing so combines two of the above non-electrophilic Nrf2 activators, physical exercise and photobiomodulation, hasn’t been a consideration.
These reviewers didn’t include human studies of sunlight’s effects. Nevermind that hospitals used to have sundecks for patients, and John Ott published relevant human and animal studies over fifty years ago.
Many studies have an undisclosed limitation in that they were performed without controlling for light. For example, knowing that mitochondria are light-activated, I don’t trust those studies’ in vivo, ex vivo, or in vitro results.
None of the 100 most recent 2025 photobiomodulation papers examined natural sunlight. Maybe it wouldn’t sell red light, green light, and blue light lasers and other products to show that people could produce the same effects themselves with sunlight at different times of the day? Would researchers damage their reputations to study a freely-available intervention, one where they don’t “do something”?