This 2018 Chinese rodent study found:
“Elevated Dnmt3a [a DNA methyltransferase] level in the dorsal dentate gyrus (dDG) of hippocampus was associated with the absence of fear renewal in an altered context after extinction training. Overexpression and knockdown of Dnmt3a in the dDG regulated the occurrence of fear renewal in a bi-directional manner.
We found that renewal of remote fear memory can be prevented, and the absence of renewal was concurrent with an elevated Dnmt3a level.
Our results indicate that Dnmt3a in the dDG is a key regulator of fear renewal after extinction, and Dnmt3a may play a critical role in controlling fear memory return and thus has therapeutic values.”
The study was a collection of five experiments investigating causes and effects of biology and behavior. The researchers used different techniques to achieve their goals. I’ve quoted extensively below to show some background and results.
“Alterations in histone acetylation and DNA methylation are involved in the formation and extinction of long-term memory. DNMTs catalyze the cytosine methylation and are required to establish and maintain genomic methylation.
Dnmt3a and Dnmt3b are de novo DNA methyltransferases. Dnmt1 is the maintenance DNA methyltransferase.
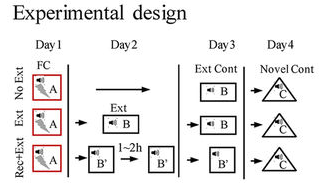
- Dnmt3a expression was elevated in the dDG after extinction training followed by a brief memory retrieval (Rec+Ext), which was associated with the absence of fear renewal when tested in an altered context.
- Increasing Dnmt3a expression in the dDG using AAV [recombinant adeno-associated virus] expression led to the prevention of fear renewal following a standard extinction training protocol.
- Knockdown of Dnmt3a in the dDG using CRISPR/Cas9 resulted in fear renewal following Rec+Ext protocol.
- Renewal of remote fear memory can be prevented using the Rec+Ext protocol.
- The absence of renewal was concurrent with an elevated Dnmt3a level.
Current exposure therapy, although effective in many patients, suffers from the inability to generalize its efficacy over time, or is limited by the potential return of adverse memory in the new/novel contexts. These limitations are caused by the context-dependent nature of extinction which is widely viewed as the biological basis of exposure therapy.
Achieving a context-independent extinction may significantly reduce fear renewal to improve the efficacy of exposure therapy. Our current study suggests that the effectiveness of these approaches, and ultimately the occurrence of fear renewal, is determined by the level of Dnmt3a after extinction training, especially in the dDG.
There are two potential mechanisms underlying extinction, one is erasure or updating of the formed memory, and the other is the formation of a new extinction memory which suppresses or competes with the existing memory in a context-dependent manner. While most studies favor the suppression mechanism in the adult, limited studies do suggest that erasure occurs in the immature animals.
We propose that if Dnmt3a level is elevated with extinction training (such as with Rec+Ext protocol), modification to the existing memory occurs and as a consequence extinction does not act as a separate mechanism or form a new memory; but if Dnmt3a level is unaltered with extinction training, a separate extinction memory is formed which acts to suppress or compete with the existing memory.”
The relevant difference between humans and lab rats is that we can ourselves individually change our responses to experiential causes of ongoing adverse effects. Standard methodologies can only apply external treatments such as exposure therapy and manipulating Dnmt3a levels.
https://www.nature.com/articles/s41598-018-23533-w “Dnmt3a in the dorsal dentate gyrus is a key regulator of fear renewal”