This 2023 rodent study wrapped together findings of the original study curated in A rejuvenation therapy and sulforaphane, and the second follow-on study mentioned in Signaling pathways and aging. I’ll start by highlighting specifics of the later study:
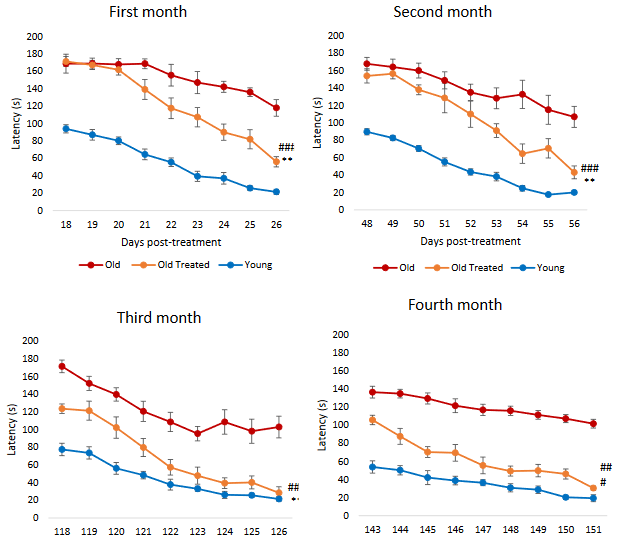
“Pronounced rejuvenation effects in male rats prompted us to conduct further confirmatory experiments. A particularly important consideration is the effectiveness of E5 with regards to sex, as sex-dependent rejuvenation by some interventions have previously been reported.
To assess E5’s applicability to both male and female Sprague Dawley rats, we studied 12 males (6 treated with E5, 6 with saline) and 12 females (6 treated with E5, 6 with saline). These rats were treated every 45 days with an injection of E5 or saline. Rats were monitored for 165 days, and blood was drawn at six time points: 0, 15, 30, 60, 150 and 165 days from the first injection.
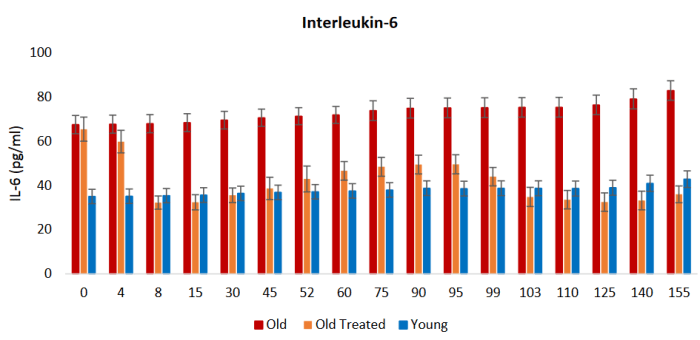
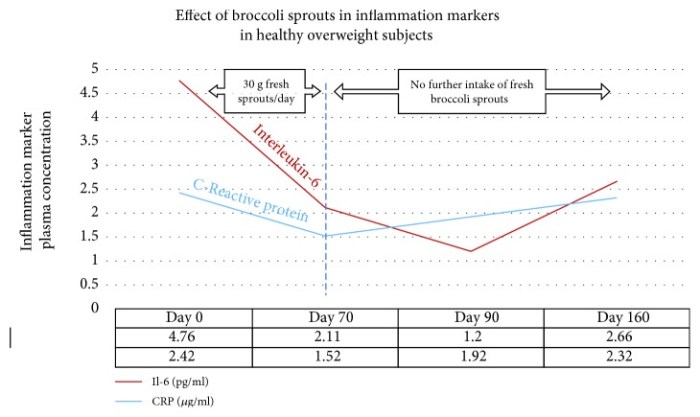
We observed highly significant improvements in TNF alpha and IL-6 levels for both males and females in the blood of E5-injected rats over that of saline controls. We also observed a substantial improvement in grip strength.
Our study shows age reversal effects in both male and female rats, but E5 is more effective in males.”
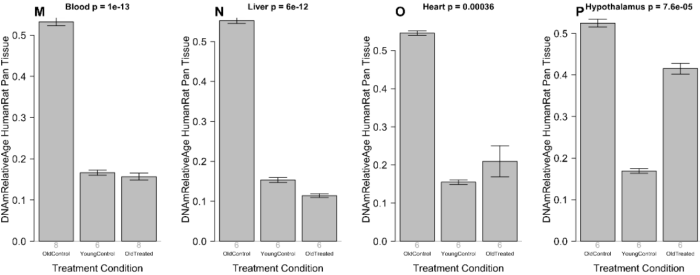
Another experimental group was started with old rats of both sexes. Using the human / rat relative clock developed in the original study, a human equivalent age to these rats at 26 months old was ((112.7 weeks / 197.6 weeks maximum rat lifespan) x 122.5 years maximum human lifespan) = 69.8 years:
“To validate our epigenetic clock results, we conducted a second set of E5 experiments with Sprague Dawley rats of both sexes. When these rats turned 26 months old, half (9 rats) received the E5 treatment while the other half (8 rats) received only the control treatment (saline injection). We analyzed methylation data from two blood draws: blood draw before treatment (baseline) and a follow up sample (15 days after the E5/saline treatment).”
Treatment measurements were affected by one female control group outlier. Panels F through J were recalculated after removing the outlier to show significant effects in both sexes:

“A) Final version of the rat clock for blood. Baseline measurement (x-axis) versus follow up measurement (15 days after treatment, y-axis). Points (rats) are colored by treatment: red=treated by E5, black=treated with saline only. Rotated grey numbers underneath each bar reports the group sizes. Each bar plot reports the mean value and one standard error.
B,D,E) Difference between follow up measurement and baseline measurement (y-axis) versus treatment status in B) all rats, D) female rats only, E) male rats only. C) is analogous to B) but uses the pan tissue clock for rats.
Panels in the second row (F,G,H,I,J) are analogous to those in the first row but the analysis omitted one control rat (corresponding to the black dot in the lower right of panel A).”
https://www.biorxiv.org/content/10.1101/2023.08.06.552148v1 “Reversal of Biological Age in Multiple Rat Organs by Young Porcine Plasma Fraction”
A description of how E5 plasma fraction was made starts on page 16 of the *.pdf file. The next E5 study will be done with dogs per July 2023 updates in blog post comments:
“On E5 our entire team is working hard towards the launch of an old Beagle dogs trial this month. We want to make them really young, healthy, happy, and jumping around like 1 and 2 year olds.
Primary endpoint is safety and toxicology to test various dose strengths and frequencies. Secondary endpoints are more than 20.
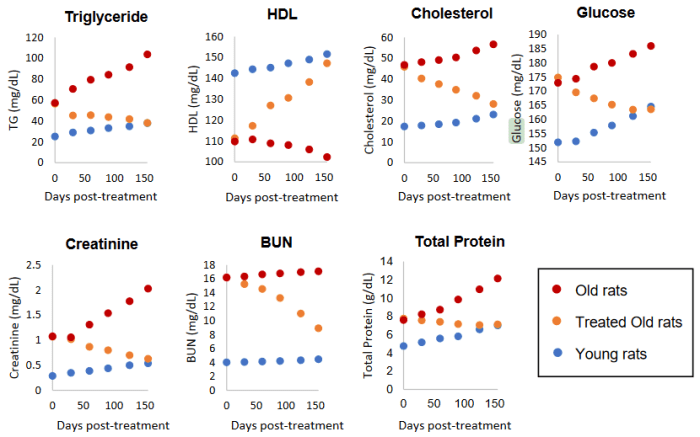
As you know, we like to test exhaustively to get a sharper perspective of what’s happening. In rat studies we tested 30 biomarkers, including functional. We are especially keen to check kidney markers.
There are two clocks for dogs we are interested in to get third party confirmation of age reversal. Horvath dog clock is ready and GlycanAge dog clock is under construction.
We are requesting all organizations that support pets and aging to financially support their project of building an accurate dog clock. Not only will it help veterinary aging research like ours, but also all the dog owners that may want to know how much improvement their dog received from treatment. Dr. Matt Kaeberlain is an advisor on their project.”





 A normal distribution
A normal distribution