
A musical analogue.

A musical analogue.
This 2021 rodent study measured sequential liver changes caused by a high-fat diet:
“Using a longitudinal mouse study of diet-induced obesity in male mice, we investigated kinetics of hepatic DNA methylation and gene expression compared to those of obesity-induction to assess if they could be causal for development of insulin resistance. We aimed to find out if these changes were reversed by massive weight loss induced by vertical sleeve gastrectomy or metformin treatment.
We identified two CpG sites within exon 1 of Fgf21 that became gradually hypomethylated upon HFD feeding. DNA demethylation started between week two and four, to become significant at week five, and significantly correlated with hepatic Fgf21 gene expression.
These DNA methylation changes preceded development of insulin resistance, and were potentially causally involved in increased Fgf21 expression and plasma levels associated with insulin resistance. This points to a key regulatory function of gene body DNA methylation, which was eventually a compensatory response to counteract the developing insulin resistance.
HFD-induced decrease in Fgf21 DNA methylation could not be reversed by vertical sleeve gastrectomy or metformin treatment. As soon as weight loss slowed down or mice started to re-gain weight, differences in DNA methylation were no longer detected compared to sham-operated mice.

As the altered DNA methylation pattern was acquired during adulthood in differentiated cells, our data emphasize that metabolic programming via DNA methylation is dynamic and not restricted to fetal development. This supports the concept that individuals can actively influence their DNA methylation patterns by lifestyle choices.
Our data indicate that DNA methylation alterations in key metabolic tissues can be acquired by an obesogenic diet, and not easily be reversed by interventions common in obese and diabetic subjects.”
https://www.sciencedirect.com/science/article/pii/S0955286321003272 “Dietary induction and reversal of obesity and insulin resistance is associated with changes in Fgf21 DNA methylation in liver of mice”
This study attempted two interventions that didn’t have desired effects. All about the betaine mentioned others that may reverse liver epigenetic changes.

This 2021 rodent study investigated sulforaphane pretreatment’s role in reducing liver injuries:
“As a double blood supply organ of the portal vein and artery, the liver is highly sensitive to ischemia, and is one of the common organs to suffer from hepatic ischemia-reperfusion injury (HI/RI). HI/RI leads to overproduction of reactive oxygen species (ROS).
- Overdoses of ROS promote reaction of lipid peroxidation and generation of the extremely aggressive oxidants nitric oxide (NO) and malondialdehyde (MDA).
- HI/RI decreased antioxidant enzyme activity of glutathione (GSH), catalase (CAT), and superoxide dismutase (SOD).
- Sulforaphane (SFN) intervention significantly decreased levels of MDA and NO by increasing activity of GSH, CAT, and SOD.
Inflammation is the most serious secondary injury experienced in HI/RI.
- Monocyte chemotactic protein 1 (MCP-1) is involved in an inflammatory reaction with regulation of monocytes, T lymphocytes, and NK cells. MCP-1 can also increase infiltration of inflammatory cells by activating NF-κB.
- Tumor necrosis factor-α (TNF-α) is a promoter of the inflammatory response. Interleukin-6 (IL-6) is an inflammatory mediator in the acute reaction period.
- SFN treatment significantly decreased HI/RI-induced expression of TNF-a, IL-6, and MCP-1.

In conclusion, SFN has a protective effect on HI/RI. The mechanism is associated with activating Nrf2/HO-1 signaling to suppress oxidative stress and inflammation.”
https://www.sciencedirect.com/science/article/abs/pii/S0966327421000794 “Sulforaphane alleviates hepatic ischemia–reperfusion injury through promoting the activation of Nrf-2/HO-1 signaling” (not freely available)
A human equivalent of this study’s 5 mg / kg sulforaphane dose was (.161 x 5 mg) x 70 kg ≈ 56 mg. For comparison, my estimated daily sulforaphane intake from microwaved sprouts is 52 mg.
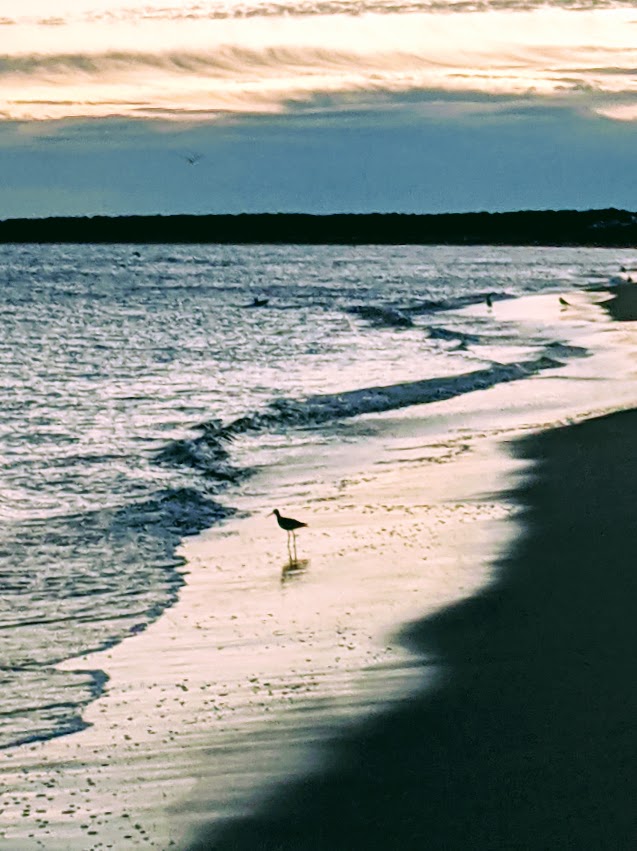

Twin 2021 in vitro studies of cruciferous sprout bioaccessibility, with the first addressing hydroxycinnamic acids and flavonols:
“The present work studies effects of physicochemical and enzymatic characteristics of gastrointestinal digestion on two major groups of phenolic compounds – flavonols and cinnamoyl derivatives – on red radish, red cabbage, broccoli, and white mustard sprouts. Effects of gastrointestinal digestion on release and stability of phenolic compounds depends on different factors, such as physicochemical traits of the food matrix, pH, temperature, or enzymatic activity.
Although initial concentrations of phenolic acids in red radish were lower than in other sprouts, their bioaccessibility after digestion was higher, followed by red cabbage, white mustard, and broccoli. Most degradation of phenolic compounds corresponded to the flavonol fraction, which was almost erased during digestion (with the exception of digestion products of broccoli sprouts, which retained around 30% of the original flavonol concentration):

Red radish sprouts exhibited the greatest bioaccessibility.
Gastric digestion prepares the food matrix for more efficient polyphenol extraction during intestinal digestion, in which the highest release and stability of these compounds takes place. Hydroxycinnamic acids reach higher concentrations than flavonols, making them tentatively more available to be absorbed at the intestinal level.”
https://www.mdpi.com/2072-6643/13/11/4140/htm “In Vitro Evidence on Bioaccessibility of Flavonols and Cinnamoyl Derivatives of Cruciferous Sprouts”
A cited predecessor used similar methods to study glucosinolate breakdown products like sulforaphane, iberin, and indole-3-carbinol:
“Significantly higher bioaccessibility of isothiocyanates (ITCs) and indoles from glucosinolates (GSLs) of red cabbage sprouts were observed. Bioaccessibility of GSLs from Brasicaceae sprouts is not exclusively associated with initial content of these compounds in plant material (almost negligible), but also with release of GSLs and ongoing breakdown reactions during gastric and intestinal phases of digestion, respectively:

The intestinal phase was the most relevant for bioaccessibility of ITCs. Aliphatic GSLs provided higher bioaccessibility of their corresponding ITCs in comparison to indolic and aromatic GSLs.”
https://www.mdpi.com/1422-0067/22/20/11046/htm “Evidence on the Bioaccessibility of Glucosinolates and Breakdown Products of Cruciferous Sprouts by Simulated In Vitro Gastrointestinal Digestion”
Gastric and intestinal simulations were instructive. But rather than depending on digestion for ITCs, I “enzymatically convert to SF before oral intake” per A follow-on study to 3-day-old broccoli sprouts have the optimal yields.
Regarding phenolic compound digestion, my focus this year has been to give my gut microbiota what they want. I expect and get reciprocity from treating them well with whole oats, broccoli-red cabbage-mustard-oat sprouts, blackberries-blueberries-strawberries, quercetin from capers, etc. polyphenols. Not to mention inulin, artichoke hearts, and yeast cell wall β-glucan. Haven’t considered sprouting red radish seeds.
Per Red cabbage effects on gut microbiota, a related research group had an in vitro system that included gut microbiota. Maybe these researchers will get together in a future study?

This 2021 human study used epigenetic clock technology to assess chronic inflammation as a driver of cognitive decline through its effects on brain structure:
“An epigenetic measure of C-reactive protein (DNAm CRP) was assembled for each participant. We found that higher inflammatory burden, indexed by DNAm CRP scores, associated with poor cognitive and neuroimaging brain health outcomes.

DNAm CRP exhibited significantly larger associations with brain structural MRI metrics (including global grey and white matter atrophy, poorer white matter microstructure, and increased white matter hyperintensity burden) than serum CRP. Given that the 7 CpGs which make up DNAm CRP score reside in inflammation and vascular-related genes, these DNAm CRP-brain MRI associations may be capturing the impact of upstream inflammatory activity above and beyond that of serum CRP levels.
Our results indicate that some cognitive domains (processing speed) may be more mediated by brain structural consequences of chronic inflammation than others (verbal memory, visuospatial ability).
Our results add to the evidence base that DNAm-based predictors of inflammation may act as a quantifiable archive of longitudinal effects of these exposures – and other unaccounted for health and genetic profiles – that serum CRP levels fail to capture. By utilising an epigenetic inflammation measure, which integrates information from multiple immune-related CpG sites, we may provide a more reliable measure of chronic inflammation and thus a more comprehensive overview of consequences of chronic inflammation on brain structure and function.”
https://n.neurology.org/content/early/2021/11/17/WNL.0000000000012997.long “DNA Methylation and Protein Markers of Chronic Inflammation and Their Associations With Brain and Cognitive Aging”
These researchers essentially negated many of their findings by acknowledging:
“Although we endeavoured to remove participants with cognition-related pathology, these were screened via self-reported diagnoses, and we may be missing undiagnosed or subclinical incident neurodegenerative pathology.”
It wasn’t sufficient to claim in the Abstract section “Participants (N = 521) were cognitively normal, around 73 years of age” then include in the Discussion section a one-sentence limitation of relying on self-reports. Everyone defends themself against current and past realities and experiences.
Hard to imagine that objective measures such as the three comprising cognitive ability weren’t better screens. But then too many 73-year-old subjects may not have been “cognitively normal” and this study wouldn’t be adequately powered?
Can humans counteract inflammation? Non-communicable diseases? Smoking? Immune system degradation? Yes. No personal-agency actions were mentioned.
Also note this study’s social norming. The above-pictured 30-year-old female was busy at work, and subsequently hoisted a cat instead of a child in later years.
Take responsibility for your own one precious life.

This 2021 human twin study used four epigenetic clocks:
“We examined the mediating role of lifestyle factors on the association between sex and biological aging in younger and older adults. The Finnish Twin Cohort (FTC) includes three large cohort studies:
- The older FTC includes twins born before 1958;
- Finntwin16 includes twins born in 1975-1979; and
- Finntwin12 includes twins born in 1983-1987.
In comparison to women, men were biologically older and, in general, they had unhealthier life habits. The effect of sex on biological aging was partly mediated by body mass index and, in older twins, by smoking. Sex was directly associated with biological aging, and the association was stronger in older twins.
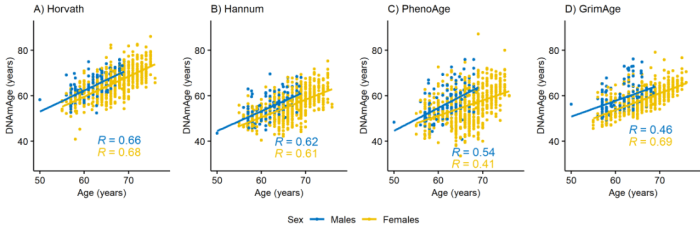
Declining smoking prevalence among men is a plausible explanation for narrowing of the difference in life expectancy between sexes. Data generated by epigenetic clocks may help in estimating effects of lifestyle and environmental factors on aging and in predicting aging in future generations.”
https://academic.oup.com/biomedgerontology/advance-article/doi/10.1093/gerona/glab337/6424421 “Do epigenetic clocks provide explanations for sex differences in lifespan? A cross-sectional twin study”
It was too much to ask of epigenetic clocks to ferret out preclinical symptoms of lifestyles and environments accelerating aging in younger twins. Levine’s Phenotypic Age clinical measurements could assess accelerated aging trajectories, but may not have been available for this study. People who are busy abusing their bodies into non-communicable diseases have plenty of other warning signs, like abdominal obesity, high blood pressure, high blood sugar, high serum triglycerides, and low serum high-density lipoprotein.
Preclinical symptoms may be reversible by individual choices that influence lifestyle and/or environment. Effective healthspan and lifespan changes measurable by epigenetic clocks are usually limited once clinical symptoms emerge, though.
Consider this rodent study’s graphic from Part 2 of Eat broccoli sprouts for your eyes:

This chart demonstrated that preventing diabetes’ negative effects on retinal function (i.e. controls) was measurably better than trying to fix subjects’ vision after onset of diabetes.
I would have liked this study to address a morbidity phase, where healthspan stops increasing but lifespan increases. That seems possible in twin studies, where one twin’s choices cause a healthspan halt compared to the other twin’s choices.
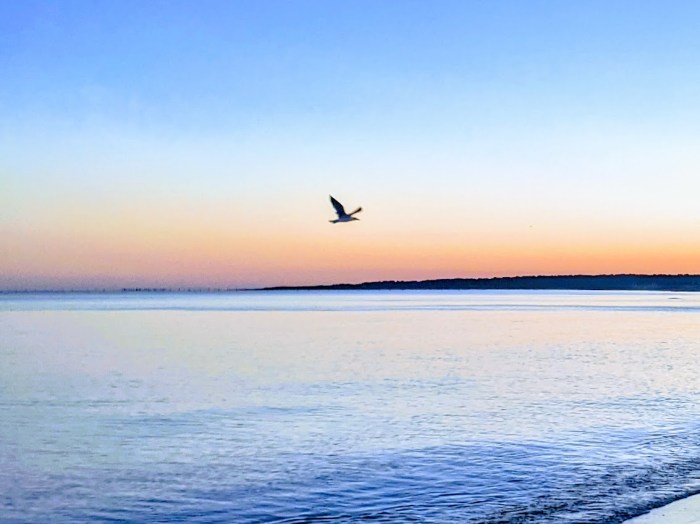

This 2021 study investigated an epigenetic treatment for bees forgetting about their hives:
“Over the last few decades, numbers of both wild and managed bee pollinators have been declining. Although reasons for this decline are under debate, it is highly likely that a combination of multiple stressors is to blame, in particular, deformed wing virus (DWV).
Histone deacetylase inhibitors (HDACi) are a class of compounds which prevent deacetylation of histones and therefore increase gene expression. The present study found that HDACi sodium butyrate (NaB) significantly increased survival and reversed the learning / memory impairment of DWV-infected bees. We demonstrated the mechanism of how epigenetic regulation can resume honeybees’ memory function.

- When bees were infected with DWV, 50% of bees died by the end of day 2 and only 10% survived to the end of day 5.
- When NaB was added to the diet prior to DWV infection, survival rate of DWV-infected bees (N/D group) remained >90% after 5 days.
- Under laboratory rearing conditions, around 30% of control bees died over a period of 5 days.
- When NaB was included in uninfected bees’ diet, less than 15% of bees died.
These results indicate that feeding bees with NaB could significantly increase survival with or without DWV infection.”
https://www.cell.com/iscience/fulltext/S2589-0042(21)01024-5 “Real-time monitoring of deformed wing virus-infected bee foraging behavior following histone deacetylase inhibitor treatment”
Interesting that these researchers didn’t attempt to eliminate either the virus cause of bee behavior or parasitical mites that carried the virus. They mainly depended on bees’ endogenous systems providing beneficial responses when stimulated.

Two 2021 papers, with the first studying bee infections:
“Fungus Nosema ceranae represents one of the primary bee infection threats worldwide, and antibiotic fumagillin is the only registered product for nosemosis disease control. Natural bioactive compounds deriving from glucosinolate–myrosinase (GSL–MYR) in Brassicaceae plants, mainly isothiocyanates (ITCs), are known for antimicrobial activity against numerous pathogens and health-protective effects in humans.
This work explored Brassica nigra [black mustard] and Eruca sativa [arugula] defatted seed meal (DSM) GSL-containing diets against natural Nosema infection in honeybees. Feeding was administered in May to mildly N. ceranae-infected colonies for four weeks at 250 g/week.
- N. ceranae abundance showed a slight but significant decrease.
- No significant effects on colony development and bee mortality were observed compared to controls.
- MYR activity was detected both in bees fed DSMs and controls.
- ITCs were found in gut tissues from bees treated with DSMs, corroborating presence of a MYR-like enzyme capable of hydrolyzing ingested GSLs.
Use of DSMs containing GSLs represents a promising alternative to fumagillin as it would overcome the problem of toxic bee product residues encountered with antibiotic treatment.”
https://www.mdpi.com/2218-273X/11/11/1657/htm “Glucosinolate Bioactivation by Apis mellifera Workers and Its Impact on Nosema ceranae Infection at the Colony Level”
A review was cited for “ITCs are GSL hydrolysis products known for their broad-spectrum biological activities against pests and soil/food-borne fungi, bacteria, and human microorganisms.”
“ITCs are efficient agents against a wide range of fungal strains. Many plant and human pathogens, as well as other fungi, were shown to be inhibited in vitro by these agents.
ITC-containing, chemically-characterized plant matrices used to test antifungal activity are summarized. The same activities were tested in pure ITCs; in fact, several well-designed studies did both approaches.
Although there is no one-size-fits-all rule to predict antifungal activity, in general, less polar compounds are usually more potent in solution, but lag behind more volatile compounds with small molecule size in vapor-phase applications. The main biochemical targets seem to be directly related to chemical reactivity, with antioxidant machinery a well-identified target.
Excellent studies on transcriptomics show general stress responses. Inhibiting production of fungal toxins is shown in many real-life applications as well.
Additional applications include:
- Inhibition of fungal growth, pathogenesis, and/or toxin production in a variety of stored plants and grains;
- Inhibition of disease on post-harvest fruits; as well as
- Increasing shelf-life of different food products.
What is more, decrease in decay is frequently accompanied by the lack of measurable changes in various quality characteristics.
ITCs’ natural origin and biodegradability make them good candidates for a wide range of possible applications. Long-term studies show effects are delivered usually without apparent side-effects to plants.”
https://www.mdpi.com/2309-608X/7/7/539/htm “Effects of Glucosinolate-Derived Isothiocyanates on Fungi: A Comprehensive Review on Direct Effects, Mechanisms, Structure-Activity Relationship Data and Possible Agricultural Applications”
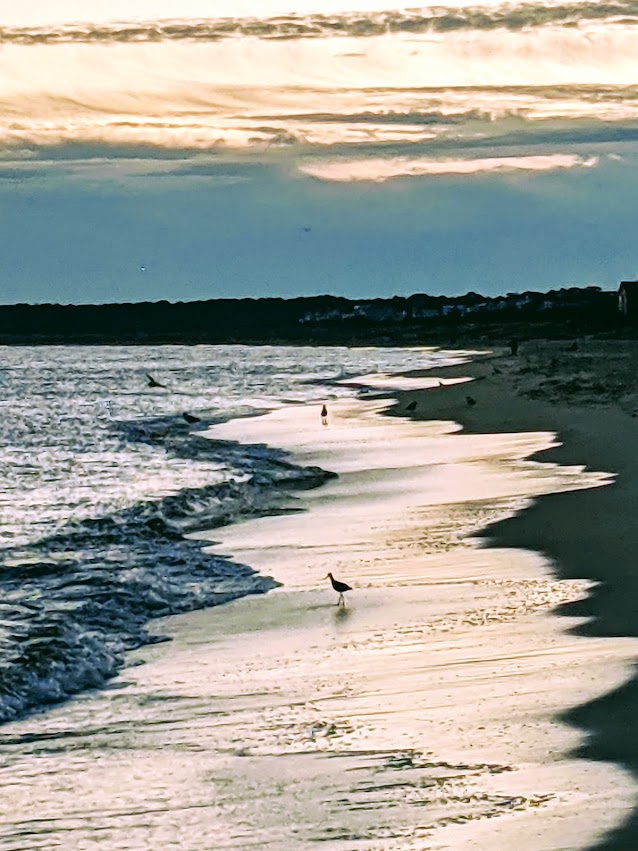

Dr. Michael Skinner coauthored a 2021 review arguing for inclusion of epigenetic transgenerational inheritance into evolutionary theory:
“Over the past 50 years, molecular technology has been used to investigate evolutionary biology. Many examples of finding no correlated genetic mutations or a low frequency of DNA sequence mutations suggest that additional mechanisms are also involved.
- Identical twins have essentially the same genetics, but generally develop discordant disease as they age.
- Only a low frequency (generally 1% or less) of individuals that have a specific disease have a correlated genetic mutation.
- Dramatic increases in disease frequency in the population cannot be explained with genetics alone.
DNA methylation, histone modifications, changes to chromatin structure, expression of non-coding RNA, and RNA methylation can directly regulate gene expression independent of DNA sequence. These different epigenetic factors do not only act independently, but integrate with each other to provide a level of epigenetic complexity to accommodate the needs of cellular development and differentiation.
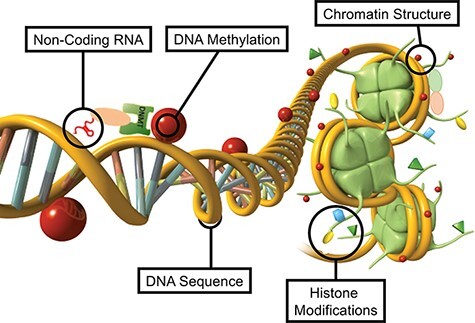
Environmental epigenetics is the primary molecular mechanism in any organism that is used to promote physiological and phenotypic alterations. Actions of environmental factors early in development can permanently program the cellular molecular function, which then impacts later life disease or phenotypes.

Integration of epigenetics and genetics contribute to a Unified Theory of Evolution that explains environmental impacts, phenotypic variation, genetic variation, and adaptation that natural selection acts on. The current review expands this proposed concept and provides a significant amount of supporting literature and experimental models to support the role of environmentally induced epigenetic transgenerational inheritance in evolution.”
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8557805/ “Role of environmentally induced epigenetic transgenerational inheritance in evolutionary biology: Unified Evolution Theory”
Organisms cited in this review’s references are similar to humans in ancestral influences and developmental influences during the first 1000 days of our lives. Humans are different in that even after all these influences, we can choose to influence our own change and individually evolve. We can also change our internal environments per Switch on your Nrf2 signaling pathway and An environmental signaling paradigm of aging.

Two 2021 papers on trained immunity, with the first a review:
“Effective memory immune responses rely on interaction between innate and adaptive immune cells. While activation of innate immunity provides the first line of defense against infections, it also primes the adaptive immune response.
Adaptive immunity can enhance antimicrobial machinery of innate cells, making them more effective at clearing pathogenic microorganisms. An additional layer of complexity adds to this network of interactions, with innate cells adopting a memory phenotype, which used to apply to only adaptive immunity. Furthermore, non-immune cells can develop some features of this memory-like phenotype.
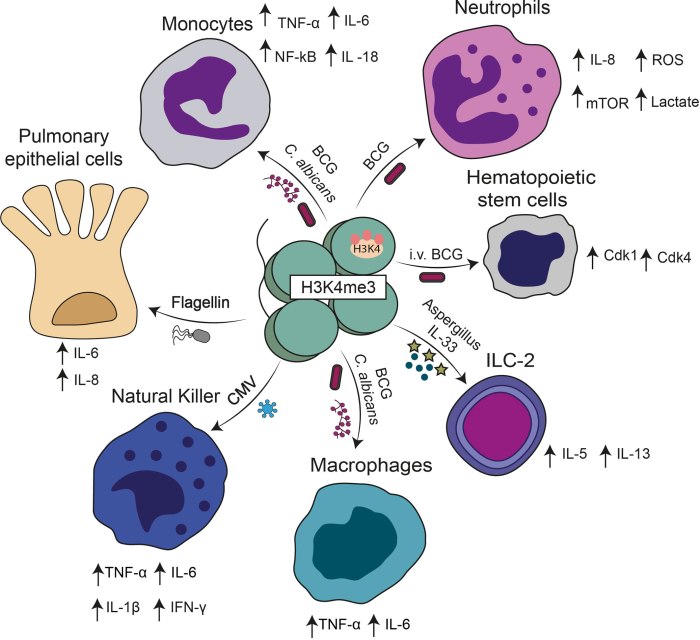
Cell subsets in which trained immunity has been described. Different stimuli including Bacillus Calmette Guerin (BCG), β-glucan, cytokines, cytomegalovirus (CMV), and bacterial components can induce a trained immunity phenotype. A common hallmark of trained immunity in these cases is H3K4me3 in promoters of genes encoding for different cytokines.
- Mechanisms Underlying Establishment of Trained Immunity
- Trained Immunity in Neutrophils
- Trained Immunity in Monocytes and Macrophages: General Features
- Metabolic Pathways Involved in Training of Monocytes and Macrophages
- Hormonal Control of Trained Immunity Responses in Monocytes and Macrophages
- Trained Immunity on Alveolar Macrophages and Involvement of Resident Cells
- Trained Immunity in NK Cells
- Trained Immunity in Innate Lymphoid Cells
- Trained Immunity on Hematopoietic Stem Cells
- Trained Immunity in Bronchial Epithelial Cells
- Trained Immunity in Skin Stem Cells
- Trained Immunity in the Gastrointestinal Tract
- Immunity Training Against Protozoan-Mediated Pathologies
- Trained Immunity in Non-Infectious Pathologies
Many gaps of knowledge remain in this field. For example, how long changes associated to trained immunity last, and if, in addition to epigenetic modulation, there are other post-translational modifications on proteins relevant for induction of trained immunity.”
https://www.frontiersin.org/articles/10.3389/fimmu.2021.745332/full “Molecular and Cellular Mechanisms Modulating Trained Immunity by Various Cell Types in Response to Pathogen Encounter”
This second paper was a human study cited for its glutathione findings as follows:
- “Plasma concentration of IL-1β from BCG-vaccinated individuals are positively associated with serum glutathione concentrations.
- Trained immunity up-regulates expression of genes involved in glutathione metabolism, suggesting an increase in glutathione synthesis and a higher glutathione recycling rate.
- Single nucleotide polymorphisms in these genes are associated with changes in pro-inflammatory cytokine production after in vitro training by β-glucan and BCG.
Enzymes whose activities are dependent on glutathione could be used as novel targets to modulate trained immunity.”

“We found a positive association between plasma glutathione concentration and ex vivo IL-1β production 90 days after BCG vaccination upon in vitro exposure to heterologous stimulus Staphylococcus aureus. Up-regulation of IL-1β production by BCG vaccination was also positively associated with circulating concentrations of other metabolites involved in glutathione metabolism, such as methionine, cysteine, glutamate, and glycine.
GSH metabolism was associated with trained immunity traits in 278 healthy individuals. Trained immunity mechanisms that are shaped by GSH metabolism remain to be further explored.”
https://www.mdpi.com/2073-4409/10/5/971/htm “Glutathione Metabolism Contributes to the Induction of Trained Immunity”
