This 2018 Baltimore cell study found:
“Based on similarities in overall methylation patterns in replicative senescence and cancers, it is hypothesized that tumor-promoting DNA methylation in cancers derives from cells escaping senescence.
We show that the tumor-associated methylation changes evolve independently of senescence and are pro-survival events with functional implications contrasting that in senescence.
In our analyses, although overall global gains and losses in DNA methylation are similar, at individual genomic regions the methylation patterns are very different for senescence versus transformation.”
https://www.sciencedirect.com/science/article/pii/S1535610818300084 “DNA Methylation Patterns Separate Senescence from Transformation Potential and Indicate Cancer Risk” (not freely available)
I hesitated to use the study’s main graphic:
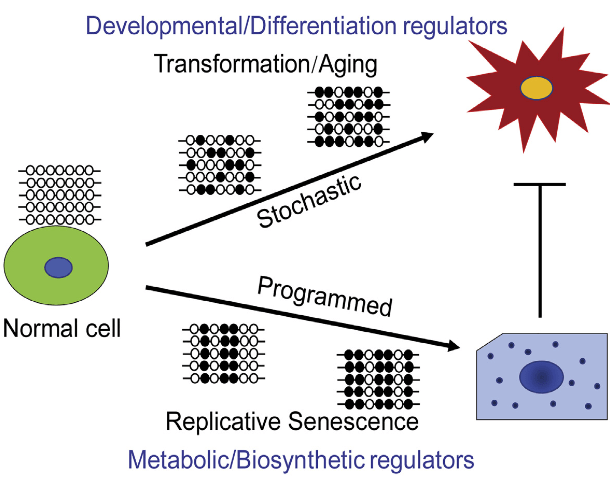 because the “Stochastic” labeling of the upper branch didn’t represent the vector’s meaning. The In Brief and the Summary sections contributed to the misrepresentation by stating:
because the “Stochastic” labeling of the upper branch didn’t represent the vector’s meaning. The In Brief and the Summary sections contributed to the misrepresentation by stating:
“transformation-associated methylation changes arise stochastically.”
which wasn’t the study’s main finding:
“Our data outlined in the above sections strongly suggest against this senescence bypass hypothesis.”
Although the experimental design and methods evoked randomness:
“Immortalization on the path to malignant transformation involves stochastic epigenetic patterns from which cells contributing to transformation may evolve.”
the graphic’s upper branch vector represented the cells’ evolutionary responses. The Significance section best characterized what the study found:
“Tumor-associated methylation changes evolve independently of senescence and are pro-survival events.”
Would anyone at John Hopkins argue, as the graphic’s upper branch labeling suggested, that cellular aging is a predominantly random process? NO!
1. Epigenetics research and evolution promoted understanding the graphic’s upper branch vector:
“Evolution is an ongoing set of iterative interactions between organisms and the environment. Directionality is introduced by the agency of organisms themselves.”
2. The current study provided another data point about the uselessness of convenient but non-etiologic, inconsequential measurements of global methylation:
“Although overall global gains and losses in DNA methylation are similar, at individual genomic regions the methylation patterns are very different.”
3. The current study was congruent with the below finding of Using an epigenetic clock to distinguish cellular aging from senescence regarding the differentiation of cellular aging from senescence:
“Cellular ageing is distinct from cellular senescence and independent of DNA damage response and telomere length.”